Ι. The Promyelocytic Leukemia (PML) and Cancer (N. Sachini, N. Papadopoulou)

Breast cancer is a heterogeneous disease (intra-/inter-tumoural heterogeneity) and remains the second leading cause of cancer deaths in women. Part of heterogeneity is proposed to be accounted by a relatively rare cancer cell subpopulation with self-renewing and aberrant differentiation properties called cancer stem cell (CSC).
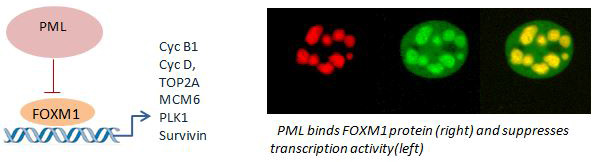
Recent studies showed that ProMyelocytic Leukemia (PML) protein regulates multiple cell functions including the stem cell properties in different tissues (hematopoietic system, nervous system and mammary gland) in normal and pathological conditions. The PML and RARα gene translocations are etiologic to Acute Promyelocytic Leukemia (APL). Historically, the generation of faithful GEM models of APL represented a fundamental step forward in demonstrating the feasibility of human cancer modeling in the mouse. PML is a multitasking regulator that controls senescence, apoptosis and stemness. It has with opposing effects on tumors, probably determined by the genetic and epigenetic cancer status under study. In the past, we have shown that PML is regulator of epigenetic transcriptional memory via its proximity to the MHC class II locus. We currently extend these by studies in tumor models in order to: a. understand the molecular signatures associated with tumor –suppressing of promoting effects by transcriptomic and proteomic (interactome, metabolomics) analysis using cell lines and patients derived data sets. b. monitor the action of inducible PML overexpression or knock down on the growth and gene expression profile, invasion and in vivo tumor growth in immunosuppressed NOD/SCID/ILRg mice. c. functionally identify PML-molecular players by rescuing its effects using CRIPR libraries guiding suitable activator or repressor Cas variants. Overall, by integrating the above via bioinformatics analysis we identify molecular targets and mediators that affect proliferation, apoptosis, DNA repair, cancer stemness and invasion the aim to develop new useful biomarkers and drug targets.
II. Polarity (M. Makkou)
Cell polarity is an evolutionarily conserved process that contributes to asymmetric cell division, migration & morphogenesis that often are disrupted in cancer.
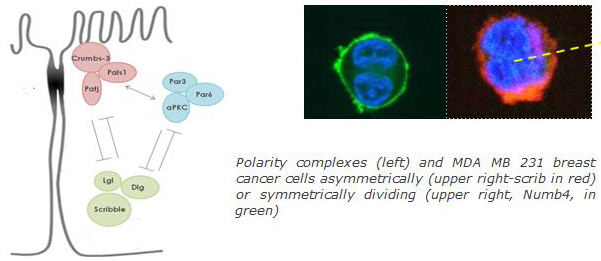
Asymmetric structure and function is a fundamental cell property that determines developmental processes and homeostasis (proliferation, apoptosis, cell-cell adhesion, asymmetric stem cell division, epithelial-mesenchymal transition). Various protein complexes and cell fate determinants such as Crumbs3-PALS1-PATJ, Par3-Par6-aPKC, Lgl-Scribble (Scrib)-Disc large (Dlg) and Numb respectively, define apical-basal, front-rear and planar cell polarity. Disruption of polarity may be due to aberrations of spindle orientation or expression and localization of cell polarity–fate determinant proteins. Such aberrations are common in various cancer types and are connected to loss of epithelial organization, cell proliferation, invasion and metastasis. Polarity genes are amplified or mutated in a large range of cancer types. Our research focuses on the effect of full length and various regions of polarity proteins in breast cancer and glioblastoma initiation and progression, applying invasion assays, sphere assays, xenografts and gene expression profiling.
III. Cancer evolution and therapy (N. Sachini, M. Makkou, M. Oraiopoulou)
Glioblastomas (GB) are highly heterogeneous, aggressive and common. Clonal and epigenetic evolution results in tumour recurrence, therapy resistance and poor prognosis in spite of recent advances. Recurrent tumors are characterized by expansion of preexisting clones as well as therapy –induced de novo mutations. Although recent sensitive techiques allow better mapping of the tumor evolution paths, they are limited in detecting minor clones. In addition tumor clonal structure has not exploited yet in guiding therapies. Addressing clonal/epigenetic evolution of resistance is crucial for developing anticipatory, precision therapeutics that will target common, recurrence associated pathways and target-specific drugs.
To approach this in GB, we combine state of the art genomic technologies and patient derived tumor xenograft (PDX).
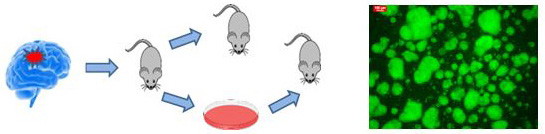
Right: From patient tumot ro cell cultuter and xenografts. Left GFP expressing glioma spheres in vitro
We apply 1. Analysis of clonal evolution upon drug pressure in vitro and in using cell barcoding (CBC) to trace and interrogate molecularly and phenotypically glioblastoma cell subsets with distinct biological properties.
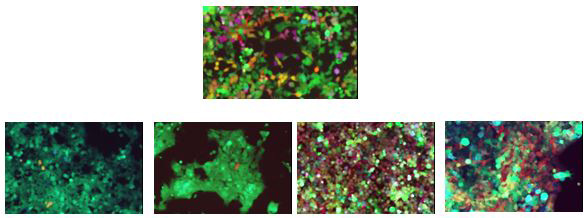
Multicolored primary patients derived glioma cells expressing variable degrees of red-green-blue fluorescent proteins in vitro (top) and after the isolation of individual metastatic foci from a xenografted mouse
2. Genetic Screens by CRISPR-dCas9 to identify susceptibility or rescue cancer related genes.
3. Development & testing of phthalimid-conjugates that lead to proteolysis (Degronimids) of known or new epigenetically acting enzyme (EZH2, DNMTs) inhibitors, in collaboration with Prof. A.Giannis (Institute of Organic Chemistry and Natural Products Chemistry, University of Leipzig).
Finally we link the above, to detailed genome structure-expression bioinformatic analysis in order to identify changes corresponding to tumor evolution and prognosis. We expect that the above in combination, will provide new valuable information in GB biology and open ways for predictions and therapies. We anticipate that the above combined studies will contribute to the understanding of GB evolution, disease recurrence and the identification of predictive biomarker in order to develop personalized therapies.




