One of the major challenges facing humankind in the 21st century is how to feed a growing population that may exceed 9.5 billion people by 2050. This needs to be done using the same or even less natural resources, because water and land are used in the increasing urbanization of the world. This implies that plant productivity needs to increase dramatically and that preventing crop losses from diseases will therefore be crucial to ensuring global food security. Plant diseases, caused by pathogenic microorganisms, reduce the yield of the world’s most important food crops, causing loss up to 40% of their annual production. One of the main reasons that plant diseases continue to cause such damage is that disease resistance introduced by plant breeding, in some cases is rapidly overcome by newly virulent pathogens. This problem is amplified by intensive crop cultivation techniques and mono-culture. Likewise, the loss of genomic diversity during the transition from wild to cultivated crop populations, enhances this problem. In order to minimize crop losses due to plant diseases we need new knowledge regarding the mechanisms underpinning microbe recognition by plants. This requires, on the one hand, to understand the biology of microbial pathogenesis and the virulence proteins they target into host cells to cause disease. From the other hand, to elucidate the mechanisms by which the resistant plants successfully defend themselves from attack.
To understand plant and animal diseases, the most important challenge is to characterize the molecular mechanisms that underlie both disease susceptibility and host immunity. Only in this way can host immunity mechanisms be predictably manipulated to provide durable disease control.
In my group we are passionate about microbes (beneficial and pathogens) and the molecular mechanisms they use to colonize their hosts. Using functional patho-genomics and patho-proteomics, we investigate the microbial strategies for host’s colonization (e.g. effector proteins, etc.), as well as the host’s innate immunity system. We use plants as host model-systems to study Molecular Host-Microbe Interactions (MHMI).
We are interested in the fundamental mechanisms regulating the 'molecular dialogs' underlying MHMI for both the host and the microbe.
Pathogens’ virulence effector proteins: Pathogenic microorganisms have evolved sophisticated strategies to exploit the attractive nutritional menu provided by their plant and animal hosts. The majority of these pathogens are highly specialized and attack only a limited number of eukaryotic host organisms. However, some strains are capable of infecting a wide range of hosts that includes both plants and animals. Plant and animal pathogens (including bacteria, fungi, oomycetes, nematodes and some insect species), translocate pathogenicity proteins (effectors), into host cells, which facilitate colonisation of susceptible hosts. Effectors have generally evolved to enable parasitism by suppressing host immunity and/or by modifying host physiology to support growth and spread of the parasite.
However, the molecular mechanisms of function and the host targets for the vast majority of the pathogens’ effectors are largely unknown. While, their activities mean that the effector proteins can be used as tools to identify important components of plant innate immunity and physiology that could potentially lead to innovative strategies for crop improvement.
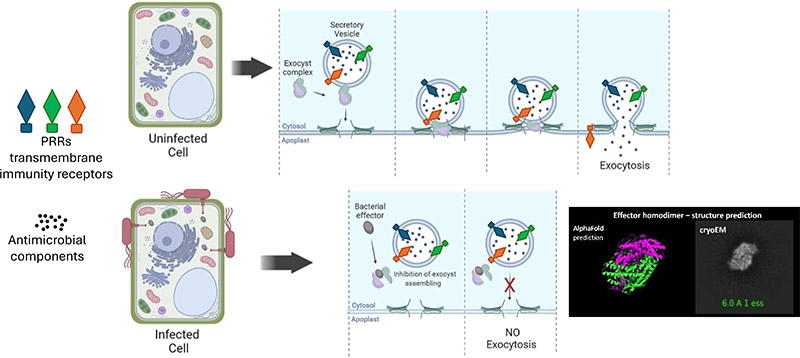
Beneficial microorganisms in plants
To ensure healthy crop production approaches, the search for alternative ecofriendly strategies - based on the EU GREEN DEAL strategy - have been proposed. The most widely accepted solution that has gained a lot of interest among scientists is the use of biological agents; mainly plant beneficial microorganisms (PBMs) that provide essential agroecosystem services within a holistic vision of enhancing farm productivity and environmental protection, according to “ONE HEALTH” strategic vision. PBMs play crucial roles in agroecological cycles fundamental for soil nutrient amelioration, crop nutrient improvement, plant tolerance to biotic and abiotic stresses, biocontrol of pests and diseases, and water uptake.
In my group we have generated a BIOBANK of more than 1500 molecularly characterized PBMs that have been isolated from CWRs (crop wild relatives) that live in extreme places (e.g. halphytes), but also from several other extreme environments (e.g. deep caves – 'Cave microbiomes'). We study the role of these microorganisms in plants' biotic and abiotic stress tolerance/resistance, but also for the identification of novel antimicrobial compounds.
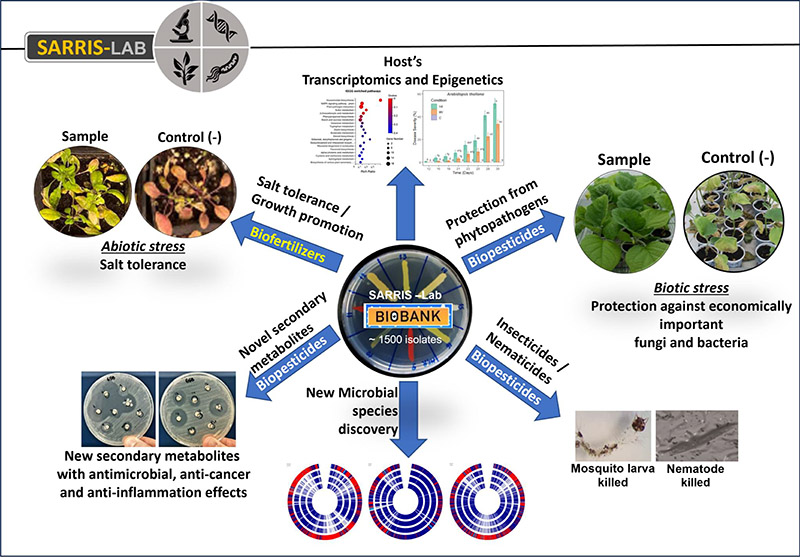
Christakis et. al., (2024) in preparation; Arapitsas et. al., (2024) in preparation
Plant innate immunity:
Another interesting research topic that we are focused in, is the study of the Plant Innate Immunity.
The innate immune system protects multicellular organisms against pathogens via specialised immune receptors that activate immunity upon detection of pathogen invasion-associated molecules. Intracellular immune receptors, called NLRs (Nucleotide-binding domain and Leucine-rich Repeat-containing), mediate innate immunity in plants and animals. In animals, NLRs typically detect conserved pathogen-associated molecular patterns (PAMPs) or damage-associated molecular patterns (DAMPs).
Plant and animal NLRs recognize pathogen-secreted pathogenicity proteins, called ‘effectors’, which promote virulence. The first characterized effectors were called ‘avirulence (AVR) proteins’ because they were recognized by an NLR encoded by a specific Resistance (R) gene allele, resulting in disease resistance, and the loss of pathogen virulence. The requirement of a specific host gene and a cognate pathogen gene for resistance led Harold Flor to postulate the ‘gene-for-gene’ hypothesis, which has acted as a framework for plant-pathogen interaction studies ever since. Upon detection of a pathogen, NLRs rapidly initiate a chain of events leading to an immune response that often culminates in programmed cell death, termed pyroptosis in animals and the hypersensitive response (HR) in plants.
Animal and plant NLRs are structurally similar: both have a nucleotide binding domain (a NACHT domain in animals and an NB-ARC domain in plants), a leucine-rich repeat domain (LRR), and an N-terminal putative signalling domain. They form part of a structural class termed signal transduction ATPases with numerous domains (STAND). Animal NLRs usually signal via either an N-terminal PYRIN domain (PYD) or caspase recruitment domain (CARD), while plant NLRs usually carry either an N-terminal TIR (Toll/Interleukin-1 receptor/Resistance protein) or CC (coiled-coil) domain.
Plants are able to defend themselves, even if (unlike mammals), they lack mobile immune cells and a somatic adaptive immune system. Plant cells are exquisitely sensitive to colonization attempts by pathogens. They rapidly perceive and respond to potential pathogen threats in a measured manner and most infections are thwarted. To understand and manipulate disease resistance in plants (and animals), we need to understand pathogens recognition by potential hosts, and how this recognition is converted into effective host defense.
A common to plants and animals defense strategy engages intracellular immune receptors. In resistant plants intracellular receptors are encoded by Resistance (R) genes. Recent genomic studies reveal that some plants carry a repertoire of up to 100-600 different R genes. These genes mostly encode intra-cellular nucleotide-binding, leucine-rich repeat NLR (Nod-like immune receptors) proteins that resemble mammalian NLR receptors. Plant NLRs recognize specific pathogen effectors, either through direct NLR/effector binding or indirectly through the perception of effector’s function or alteration of a specific host process. R proteins activate a strong immune response known as “Effector-Triggered Immunity” (ETI). ETI in plants often leads to programmed hypersensitive cell death response (HR), which prevents further pathogen growth and spread to neighboring tissues and other plants. These immune mechanisms have been successfully described in what has become known as the “zigzag” model. However, how perception leads to defense activation and HR induction is largely unknown. This constitutes a general problem for the predictive manipulation of immunity and in immune receptors function.
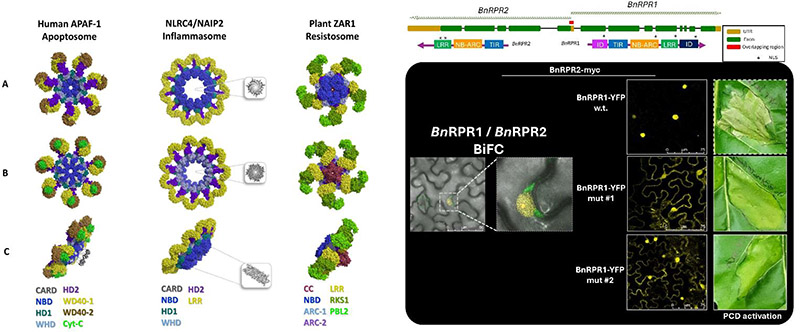
Current Aims
To understand animal and plant colonization by microbes (beneficial and pathogenic), the most important challenge is to characterize the molecular mechanisms and the macromolecular structures that underlie host immunity, as well as microbial virulence. However, the exact role(s) and the subcellular targets of microbes’ virulence components remain an important open question in MHPI field, which has only partially been elucidated.
In both plants and animals, NLR immunity receptors’ activation leads to a localized Programmed Cell Death (PCD). Recently we -and our collaborators- discovered that certain plant NLRs include additional domains to their canonical structure. We reported that these integrated protein-domains (IDs) act as “decoys” for pathogens. We found the origin of these IDs through the duplication of the effectors’ original virulence targets.
My group interests how plants resist disease using NLR-IDs, and in how resistance is overcome by pathogens.
What are the effectors’ targets (in susceptible host-cell) and how their perception by NLR-IDs activates defense, is a crucial step in our in-depth understanding of host defense mechanisms, as well as, the microbial virulence.
Additionally, in my group, we investigate the role and potential use (as Biopesticides and Biofertilizers) of the cultivated microbiome from crop wild relatives (CWR) that live in extreme environments. We isolate new and known microbes that reveal interesting features regarding their ability to produce antimicrobials (antibacterial and antifungal compounds) against pathogens of clinical and agricultural interest, as well as other interesting features (e.g. anticancer and anti-inflammatory compounds).




